Statistics is the foundational language of scientific inquiry. Artificial intelligence (AI) has rapidly emerged as a transformative technology that reshapes how knowledge is represented and discoveries are made. Interest continues to grow in harnessing the power of AI to advance scientific research. Thoughtful integration of AI and broader machine learning tools within biostatistics offers tremendous potential for innovation in both research and education. At Michigan Biostatistics, research teams play a critical role in exploring and addressing challenges at the intersection of statistics and AI—ensuring that the development and application of powerful AI technologies are both rigorous and responsible across major scientific domains. The following highlights a growing range of AI-related research areas within Michigan Biostatistics.
![]()
Foundations of Statistical and Interpretable AI
Advancing the theoretical underpinnings of trustworthy AI through rigorous statistical modeling, causal inference, and uncertainty quantification. This area bridges traditional Bayesian inference and modern machine learning, ensuring AI methods remain interpretable, robust, and generalizable.
Faculty: Erin Craig, PhD
AI models are transforming how we learn from and interpret data. What makes them so successful, and how can we distill their strengths into simpler, sparse, interpretable methods?
Example: Pretraining and the lasso
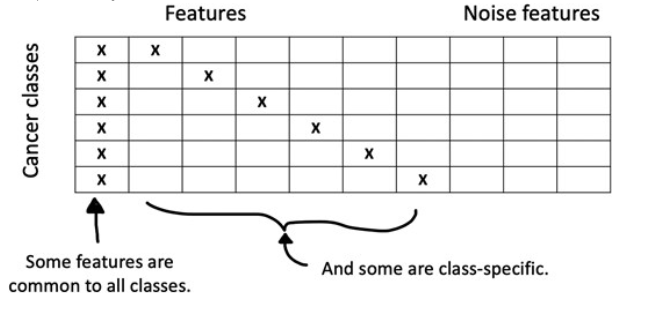
Software: R package for lasso pretraining
Faculty: Jian Kang, PhD
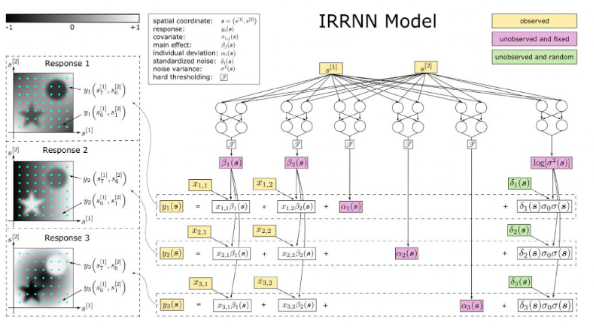
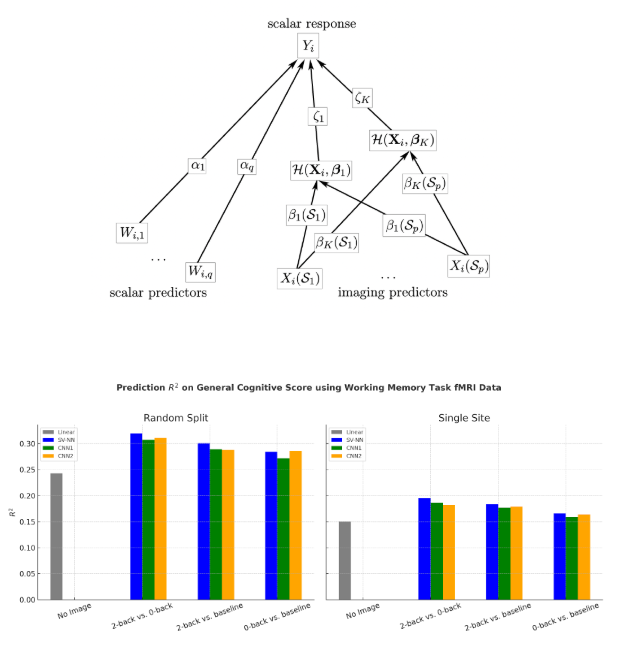
Daiwei Zhang, Lexin Li, Chandra Sripada, Jian Kang, JRSS-B (2023)
Develops IRRNN, a hybrid model integrating Spatially Varying Coefficient Models (SVCM)
with deep networks for voxel-wise association inference. Applies IRRNN to ABCD and
HCP fMRI datasets, identifying fronto-parietal and dorsal attention networks linked
to working memory performance.
Software package on GitHub
Faculty: Jian Kang, PhD
Ben Wu, Keru Wu, Jian Kang, Journal of Machine Learning Research (2025)
Introduces the SV-NN prior, a single-layer Bayesian neural network with soft-thresholded Gaussian process weights. Achieves interpretable spatial feature selection and region importance mapping in high-dimensional fMRI tasks.
Faculty: Peter Song, PhD
Professor Song has been pursuing leveraging AI and Data Science Analytics to investigate various environmental toxic agents and social stressors, which are naturally of weak signals. His lab has developed statistical learning algorithms and software to address significant challenges in exposure mixture approaches and high-dimensional mediation analyses.
Selected Publications
- Zhou, Y and Song, PXK (2025). Synergistic self-learning approach to establishing personal nutrition intervention schemes from multiple benefit outcomes in a calcium supplementation trial. Journal of Royal Statistical Society Series C 74(4):925-945.
- Wang, W, Wu, S, Zhu, Z, Zhou, L and Song, PXK* (2024). Supervised homogeneity fusion: A combinatorial approach. Annals of Statistics 52(1), 285-310.
- He, Y, Song, PXK and Xu, G (2024). Adaptive bootstrap tests for composite null hypotheses in the mediation pathway analysis. Journal of the Royal Statistical Society Series B 86, 411-434.
- Zhou, Y and Song, PXK (2023). Longitudinal self-learning of individualized treatment rules in a nutrient supplementation trial with missing data. Statistics in Medicine 42, 3032-3049.
- Hao, W and Song, PXK* (2023). A simultaneous likelihood test for joint mediation effects of multiple mediators. Statistica Sinica 33, 2305-2326. (This paper received Young Investigator Award from the ASA Section on Statistics in Epidemiology)
 Artificial Intelligence for Electronic Health Records and Clinical Decision Support
Artificial Intelligence for Electronic Health Records and Clinical Decision Support
Leveraging AI and natural language models to extract insights from electronic health records, clinical notes, and longitudinal patient data. Faculty develop reinforcement-learning frameworks for individualized treatment, adaptive trials, and real-time decision support.
Faculty: Donglin Zeng, PhD
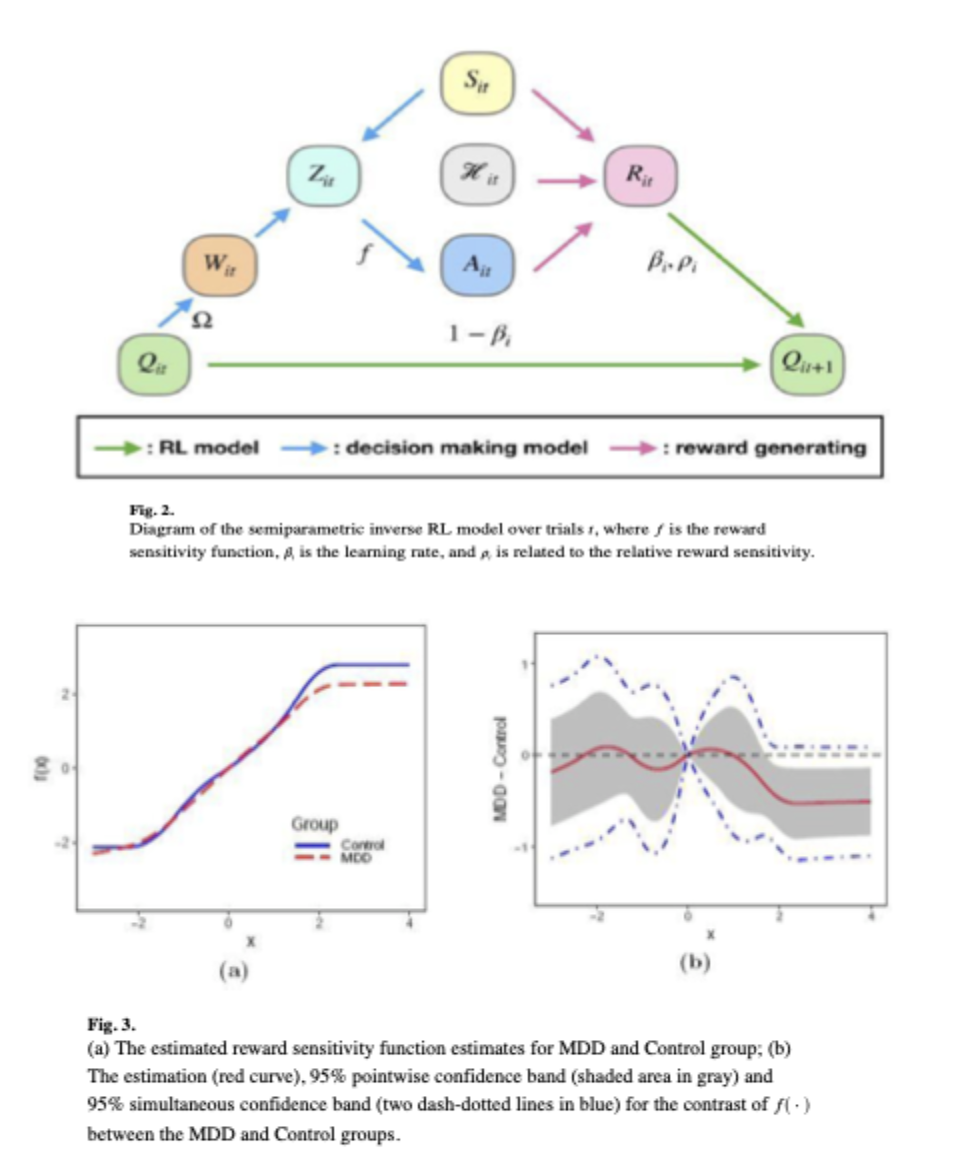
Guo, X., Zeng, D., and Wang, Y. (2024). A Semiparametric Inverse Reinforcement Learning
Approach to Characterize Decision Making for Mental Disorders. Journal of the American
Statistical Association, 119, 27-38.
Major depressive disorder (MDD) is a leading cause of disability-adjusted life years and is associated with abnormal reward processing. It remains unclear whether these abnormalities arise from reduced reward sensitivity or impaired learning. Drawing on data from the EMBARC study’s probabilistic reward task, this project proposes a semiparametric inverse reinforcement learning (RL) approach to analyze MDD patients’ reward-based decision-making. The model updates decisions via reward prediction errors weighted by individual learning rates, and models reward sensitivity as a nonlinear, nondecreasing function using I-spline estimation and joint likelihood maximization. Applying to EMBARC, we find similar learning rates but distinct, nonlinear reward sensitivity functions between MDD and control groups. These functions correlate with brain activity in negative affect circuitry during an emotional conflict task.
Faculty: Donglin Zeng, PhD
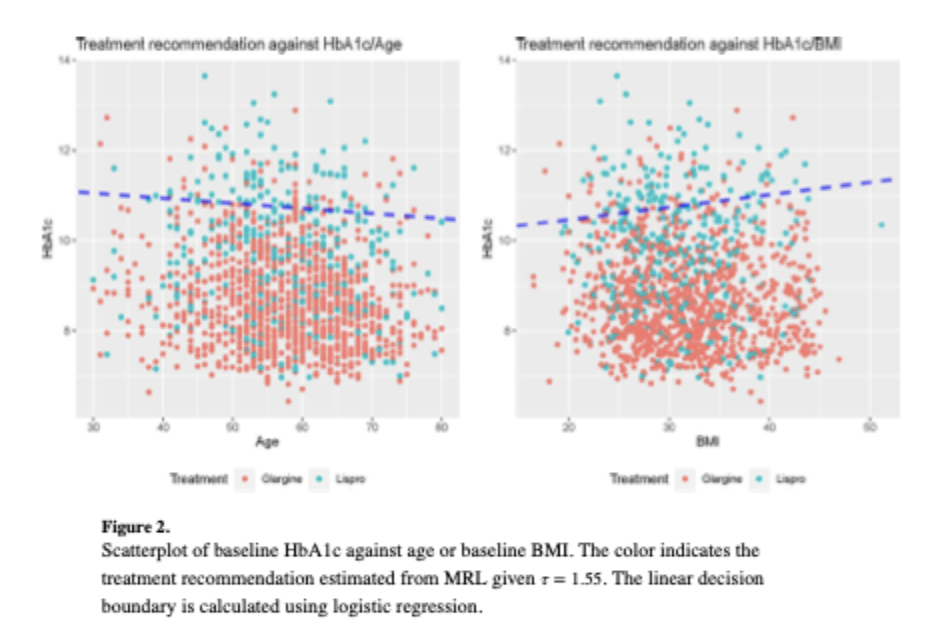
- Liu, M., Wang, Y., Fu, H., and Zeng, D. (2024). Learning Optimal Dynamic Treatment Regime Subject to Stagewise Risk Controls. Journal of Machine Learning Research, 25, 1-64.
- Liu, M., Wang, Y., Fu, H., and Zeng, D. (2024). Controlling Cumulative Adverse Risk in Learning Optimal Dynamic Treatment Regimens. Journal of the American Statistical Association, 119, 2622-2633.
Dynamic treatment regimens (DTRs) are essential for personalized medicine, particularly in conditions like cancer and type 2 diabetes, where aggressive treatments may improve efficacy but increase risk. Few existing methods estimate DTRs while balancing cumulative benefit and risk. This project develops a statistical learning framework to identify optimal DTRs that maximize outcomes and keep either short-term or cumulative risk below a set threshold. Application to a two-stage clinical trial for T2D demonstrate the method’s effectiveness to improve the control of glucose levels but effectively minimize the risk of hyperglycemia or weight gain for T2D patients.
Faculty: Donglin Zeng, PhD
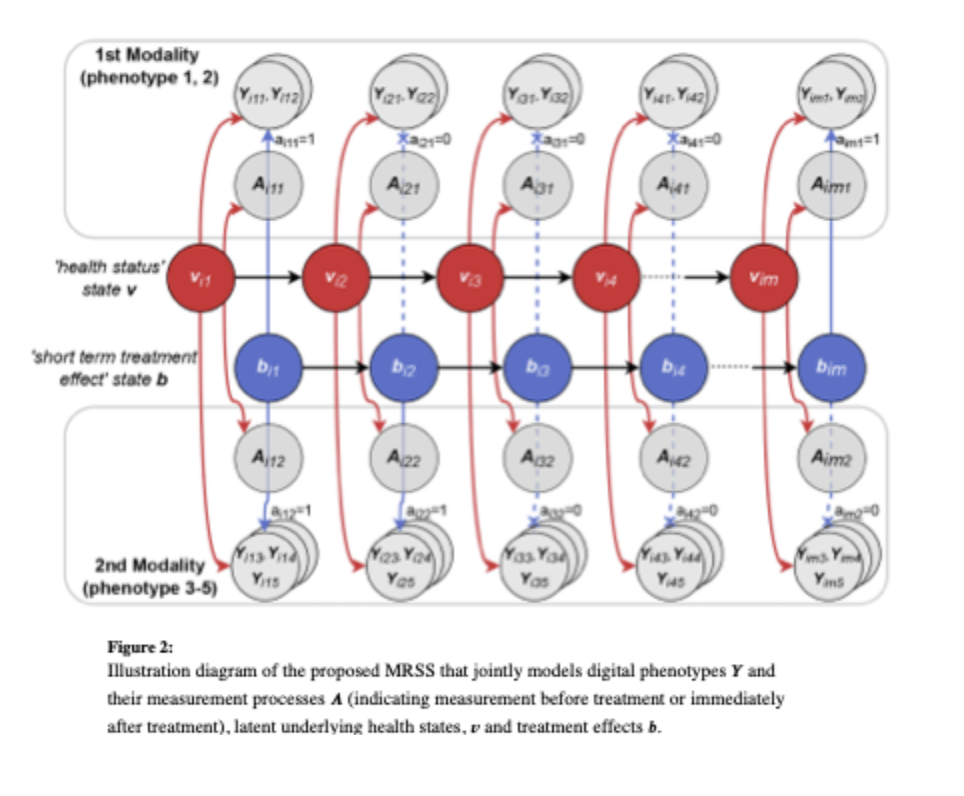
Xu, T, Chen, Y, Zeng, D., and Wang , Y. (2023). Mixed-Response State-Space Model for Analyzing Multi-Dimensional Digital Phenotypes. Journal of the American Statistical Association, 544, 2288-2300.
Digital technologies can provide frequent, objective, real-world digital phenotypes, but modeling these data is challenging due to confounding and variability from environmental factors and measurement noise. For example, signals on patients’ underlying health status and treatment effects are mixed with variation due to the living environment and measurement noises. Motivated by a Parkinson’s disease mobile health study, this project proposes a mixed-response state-space (MRSS) model to jointly analyze multidimensional digital phenotypes through latent state time series, capturing dynamic health status and treatment effects. The method uses Kalman filtering for Gaussian data and importance sampling with Laplace approximation for non-Gaussian data. The PD mobile health application show MRSS’s effectiveness in remote, real-time digital phenotype analysis.
Faculty: Peter Song, PhD
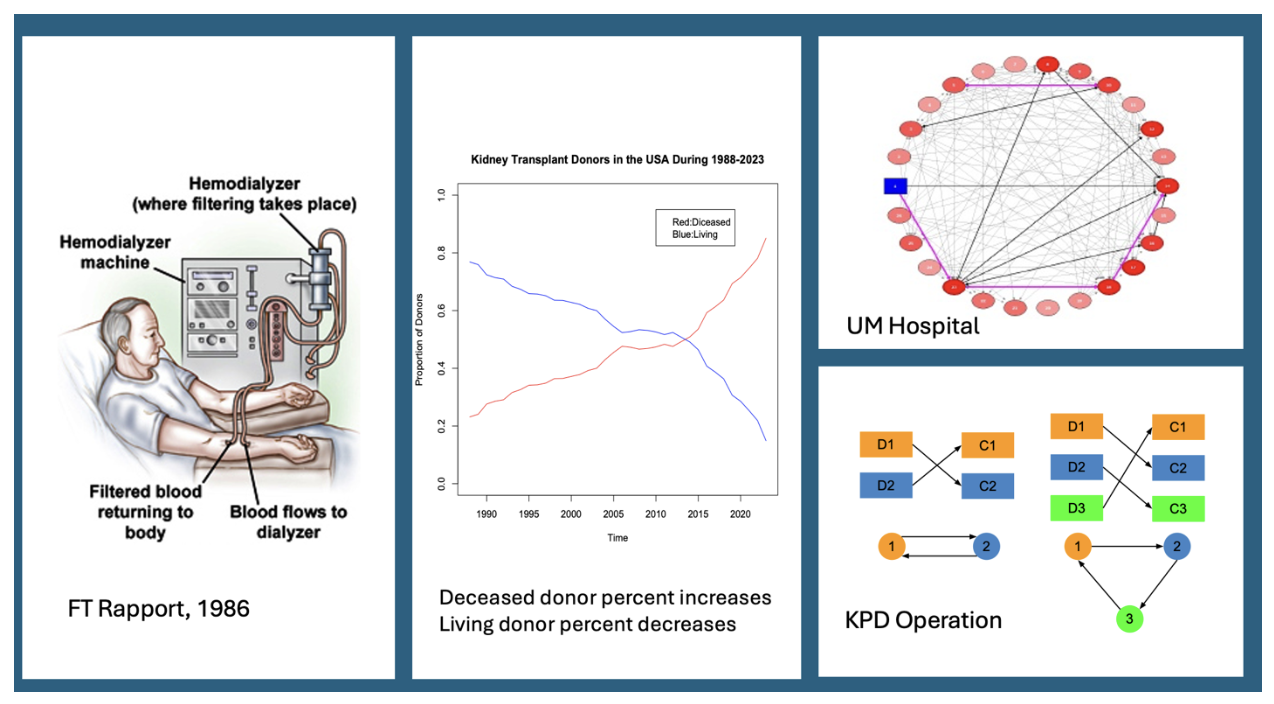
In a series of publications Professor Song and his trainees have developed different optimal organ matching strategies to match living donors with recipients in the US kidney paired donation programs in that novel strategies are evaluated by comprehensive micro simulation models that generate synthetic donors and patients with end-stage renal disease. This research program has been funded by both NIH and NSF funding agencies.
Selected Publications:
-
Wang, W, Leichtman, AB, Michael A. Rees, MA, Song, PXK, Ashby, VB, Tempie Shearon, T and Kalbfleisch, JD (2022). Kidney Paired Donation Chains Initiated by Deceased Donors. Kidney International Reports 7(6) ,1278-1288. Media coverage https://www.healio.com/news/nephrology/20220404/deceased-donor-chain-initiating-kidneys-might-increase-annual-transplants.
-
Wang, W, Rees, M, Leichtman, A, Song, PXK, Bray, M, Ashby, V, Shearon, T, Whiteman, A and Kalbfleisch, JD (2021). Deceased donors as non-directed donors in kidney paired donation. American Journal of Transplantation 21, 103-113.
-
Wang, W., Bray, M., Song, P.X.K. and Kalbfleisch, J.D. (2019). An efficient algorithm to enumerate sets with fallbacks in a kidney paired donation program. Operations Research in Health Care 20, 45-55.
-
Bray, M, Wang, W, Rees, MA, Song, PXK, Leichtmane, AB, Ashby, VA and Kalbfleisch, JD (2019). KPDGUI: An interactive application for optimization and management of a virtual kidney paired donation program. Computers in Biology and Medicine 108, 345-353.
-
Bray, M, Wang, W, Song, PXK and Kalbfleisch, JD (2018). Valuing sets of potential transplants in a kidney paired donation network. Statistics in Biosciences 10, 255-279.
-
Wang, W, Bray, M, Song, PXK and Kalbfleisch, JD (2017). A look-ahead strategy for non-directed donors in kidney paired donation. Statistics in Biosciences 9, 453-469.
-
Bray, M, Wang, W, Song, PXK, Leichtman, AB, Rees, MA, Ashby, VB, Eikstadt, R, Goulding, A and Kalbfleisch, JD (2015). Planning for uncertainty and fallbacks can increase the number of transplants in a kidney paired donation program. American Journal of Transplantation 15, 2636-2645.
-
Li, Y, Song, PXK, Zhou, Y, Leichtman, AB, Rees, MA and Kalbfleisch, JD (2014). Optimal decisions for organ exchanges in a kidney paired donation program. Statistics in Biosciences 6, 84-104.
-
Cheng, D, Song, PXK and Liu, Z (2014). Kidney paired donation system. Chinese Journal of Nephrology, Dialysis & Transplantation 23(4), 385-289.
-
Chen, Y, Li, Y, Kalbfleisch, JD, Zhou, Y, Leichtman, A and Song, PXK* (2012). Graph-based optimization algorithm and software on kidney exchanges. IEEE Transactions on Biomedical Engineering 59(7), 1985-1991.
-
Chen, Y, Kalbfleisch, JD, Li, Y, Song, PXK and Zhou, Y (2011). Computerized platform for optimal organ allocations in kidney exchanges. BIOCOMP2011 Conference (acceptance rate 21%).
Faculty: Peter Song, PhD
Research Theme: Foundations of Machine Learning in Multicenter Clinical Studies
A distributed analysis is conducted under a colectively agreed analysis protocol as well as constant communication/consultation across study sites. Data privacy is protected since it only requires sharing of summary statistics. No loss of statistical power in comparison to the analysis with the traditional centralized database. Implemented for (Weighted) Generalized Linear and time-to-event outcomes.
Software package on GitHub: https://github.com/CollaborativeInference
Selected Publications:
- Hu, M, Shi, X, Gong, Z and Song, PXK* (2025). Collaborative inference for accelerated failure time model using clinical center-level summary statistics. Statistics in Medicine (to appear).
- Hu, M, Shi, X and Song, PXK* (2024). Collaborative inference for treatment effect with distributed data-sharing management in multicenter studies. Statistics in Medicine 43, 2043-2297 DOI:10.1002/sim.10068
- Zhou, L, She, X and Song, PXK* (2023). Distributed empirical likelihood approach to integrating unbalanced datasets. Statistica Sinica 33, 2209-2231.
- Hector, E and Song, PXK (2022). Joint integrative analysis of multiple data sources with correlated vector outcomes. Annals of Applied Statistics 6(3), 1700-1717.
- Hector, EC and Song, PXK* (2021). A distributed and integrated method of moments for high-dimensional correlated data analysis [This paper received the 2017 ENAR John Van Ryzin Award]. Journal of the American Statistical Association 116 (534), 805-818.
 Artificial Intelligence for Genomics and Multi-Omics Integration
Artificial Intelligence for Genomics and Multi-Omics Integration
Designing scalable, interpretable AI tools to analyze high-dimensional genomic, transcriptomic, and spatial-omics data. Research spans statistical deep learning, graph-based representations, and federated inference across multi-site biobanks (All of US, UK Biobank, TCGA).
Faculty: Erin Craig, PhD
B and T cell receptors are protein complexes, often represented as strings of amino acids, all with different lengths. In reality, however, they are three-dimensional, and their configurations are largely unknown. So, how should we represent them in a statistical or machine learning model? Embeddings from large protein language models can be useful representations of B/T cell receptors, but the largest signal they capture is information that is already known to scientists. To better leverage LLMs, we ask: how can we improve protein language models for immune-specific contexts? How can we use them most effectively, and is it possible to interpret their learned features in biologically meaningful ways?
Example: Disease diagnostics using machine learning of B cell and T cell receptor sequences
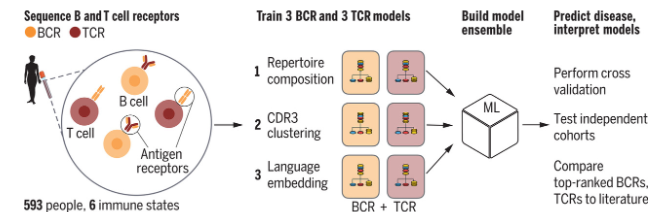
Software: Python code
 Artificial Intelligence for Imaging and Biomedical Signals
Artificial Intelligence for Imaging and Biomedical Signals
Developing AI and machine learning methods for neuroimaging, digital pathology, brain-computer interface, and other biomedical signals. Faculty innovate in spatial and temporal modeling, generative representations, and interpretable AI to map brain structure, function, and disease progression.
Faculty: Jian Kang, PhD
Tianwen Ma, Jane E. Huggins, Jian Kang, JASA Applications & Case Studies (2025)
Introduces a hierarchical Bayesian Signal Matching (BSM) model to transfer information across EEG participants, reducing calibration time for P300 BCIs. Shows strong cross-subject performance on ALS datasets and simulated EEG.
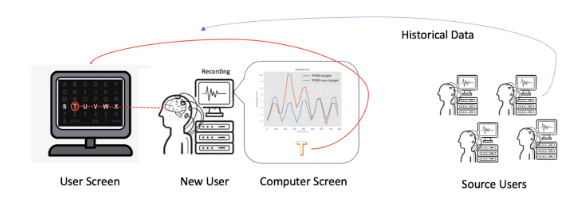
Faculty: Jian Kang, PhD
Bangyao Zhao, Yixin Wang, Jane E. Huggins, Jian Kang, Annals of Applied Statistics
(2025)
Combines Bayesian model-based RL with an actor–critic design to dynamically adjust stimulus presentation and early stopping. Improves BCI communication speed while reducing user fatigue.
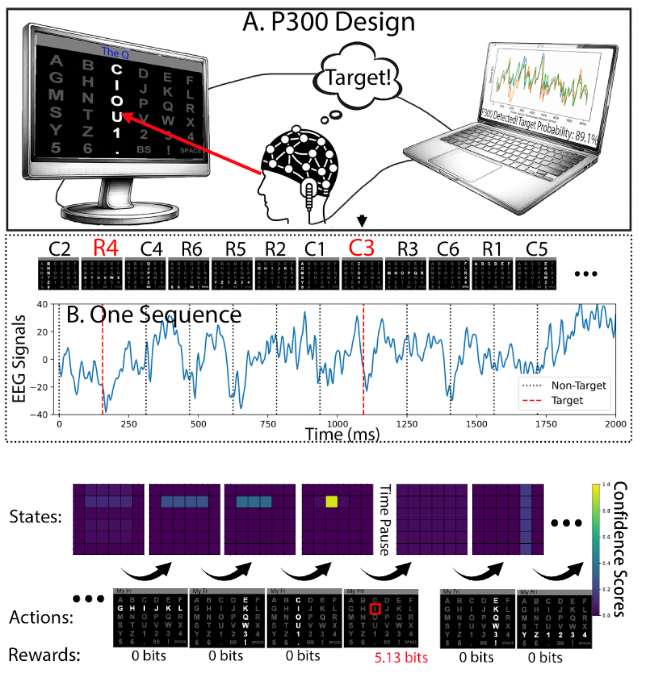
Faculty: Jian Kang, PhD
Guoxuan Ma, Bangyao Zhao, Hasan Abu-Amara, Jian Kang, Annals of Applied Statistics
(2025)
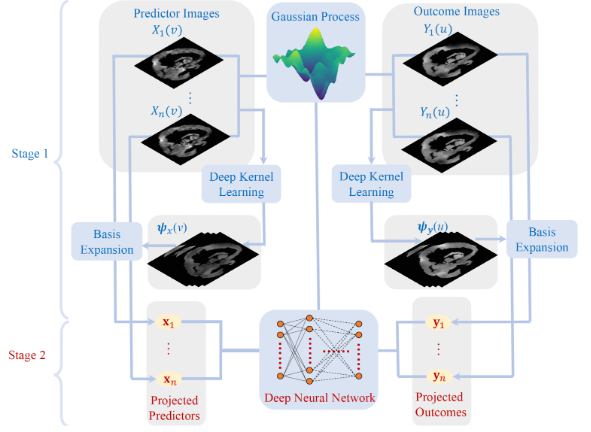
Proposes BIRD-GP, a hierarchical Bayesian Deep Kernel Learning Gaussian Process model combining neural feature learning with Gaussian process inference. Enables nonlinear cross-modal prediction between resting-state and task fMRI maps (HCP data).
 Artificial Intelligence for Mobile and Digital Health
Artificial Intelligence for Mobile and Digital Health
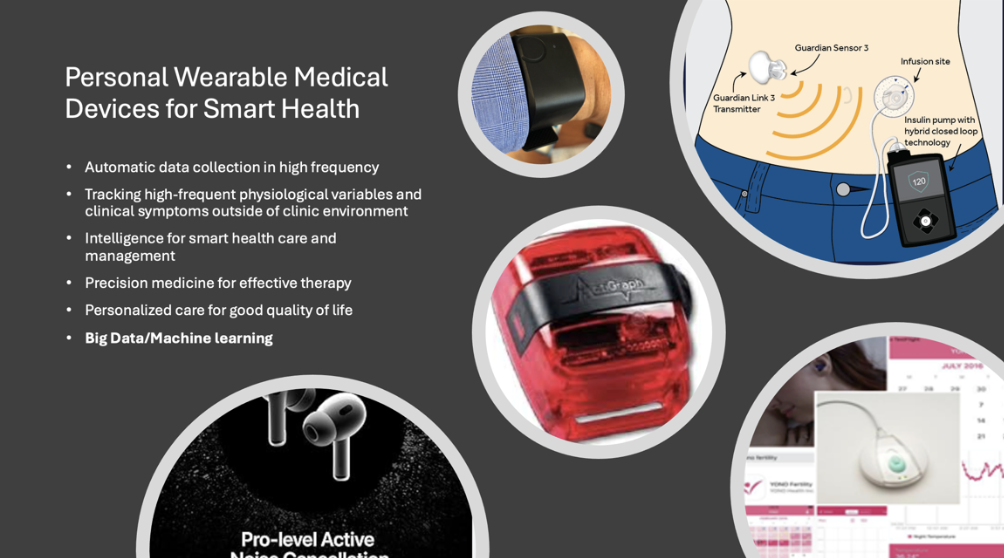
Harnessing AI, wearable sensors, and mobile data streams to understand human behavior, monitor health in real time, and enable just-in-time adaptive interventions. This theme unites statistical learning, signal processing, and reinforcement learning for personalized digital health solutions.
Faculty: Peter Song, PhD
Professor Song and his trainees have studied various wearable devices to derive digital features using AI and deep learning algorithms and evaluate influences of these digital markers on various health outcomes, such as sleep health, reproductive health, viral infection, diabetics, noise pollution, and cognitive functions. They have analyzed time-frequency time series data of physiological signals collected by various sensors.
Selected Publications:
- Banker, MM, Zhang, L and Song, PXK* (2024). Regularized scalar-on-function regression analysis to assess functional association of critical physical activity window with biological age. Annals of Applied Statistics 18(4): 2730-2752.
- Banker, M and Song, PXK* (2023). Supervised learning of physical activity features from functional accelerometer data. IEEE Journal of Biomedical and Health Informatics 27(12), 5710-5721.
- Shi, L, Wank, M, Chen, Y, Wang, Y, Hector, EC and Song, PXK* (2022). Sleep classification with artificial synthetic imaging data using convolutional neural networks. IEEE Journal of Biomedical and Health Informatics 27, 421- 432.
- Sabeti, E, Sehong Oh, S, Song, PXK and Hero, A (2022). A pattern dictionary method for anomaly detection. Entropy 24(8), 1095.
- Baek, J, Banker, M, Jansen, EC, She, X, Peterson, KE, Pitchford, EA and Song, PXK* (2021). An efficient segmentation algorithm to estimate sleep duration from actigraphy data. Statistics in Biosciences 13, 563-583. https://doi.org/10.1007/s12561-021-09309-3 (received the 2022 Statistics in Biosciences Best Paper Award).
- She, X, Zhai, Y, Henao, R, Woods, CW, Chiu, C, Ginsburg, GS, Song, PXK and Hero, AO (2021). Adaptive multi-channel event segmentation and feature extraction for monitoring health outcomes. IEEE Transactions on Biomedical Engineering 68(8), 2377-2388.
- Sabeti, E, Song, PXK and Hero, A (2020). Pattern-Based Analysis of Time Series: Estimation. Referred and accepted by The 2020 IEEE International Symposium on Information Theory, Los Angeles (acceptance rate TBA).
- Lou, L, She, X, Cao, J, Zhang, Y, Li, Y and Song, PXK* (2020). Detection and prediction of ovulation from body temperature measured by an in-ear wearable thermometer. IEEE Transactions on Biomedical Engineering 67(2), 512-522.